2026 Calibration
Lume 1.2 – Method Calibration
Paired calibration of the Lume 1.2 sensor against two EPA-approved reference methods — Colilert (IDEXX defined-substrate, MPN) and membrane filtration (MF, CFU) — and the Aquagenx Compartment Bag Test (CBT, MPN) used in international monitoring, for E. coli and total coliform quantification.
Field Validation
—Field Records
—Sensor Sites
—Latest Sample
Lab Validation
—Lab Records
—Sensors Tested
Models Running on Production Dashboards
The customer dashboards (Boulder, Denver, DC, future deployments) all load a single shared model module at /js/ecoli-model.js, served from shared/ecoli-model.js in the virridy-lume-summary repo. The two models below are what those dashboards actually evaluate against incoming sensor readings; everything else on this page is calibration evidence supporting them.
Continuous prediction
Temp + turbidity corrected single-predictor
mon2_corrected = mon2 · exp(−ρ·(T−20)) · exp(−k·NTU)
log₁₀(cfu) = b₀ + FE[barcode] + β·mon2_corrected
Fit on post-burn-in field data (n=24, 4 sensors). ρ = −0.0792, k = +0.00193. NTU derived from signal_per_spad_kcps. Sensor FE applied for calibration sensors; new deployments fall back to the reference intercept.
Field R² = 0.68
LOO R² = 0.44
RMSE = 0.45 log₁₀(MPN)
Calibration: ECOLI_CFU_COEFS
Binary alert ≥126 CFU/100 mL
3-feature logistic
P(≥126) ~ mon2 + floor_temp + tof_mean. Lab-fit logistic regression on raw features (no correction baked in). Threshold selected by Youden’s J on LOO-CV probabilities.
Lab AUROC = 0.85
Sensitivity = 1.00
Specificity = 0.79
Calibration: ECOLI_COEFS
Field Calibration Data
Live data from the mWater Lume 1.2 – 2026 Validation Data datagrid. Each water sample collection event is paired with a reference enumeration; the Method column distinguishes which reference was used — Colilert (IDEXX defined-substrate MPN), membrane filtration (MF, CFU), or compartment bag tests (CBT, MPN). The Use in Calibration column flags rows that are unusable because no /diagnostics record (water temperature, required by the CFU regression) was streaming within ±20 min of the sample. All date/time columns are displayed in UTC. Times are corrected from the mWater-stored value using the Timezone Entered column: mWater records times using the data-entry device’s local clock (Boulder, MDT = UTC−6); for samples collected in a different timezone the stored time is adjusted accordingly.
Boulder Creek E. coli Distribution — All Colilert Grabs
All unique Colilert grab samples collected at Boulder Creek sites (n = 39, deduplicated). Values in CFU/100 mL. EPA single-sample recreational threshold: 126 CFU/100 mL.
| Site |
n |
Min |
Median |
Max |
≥126 CFU |
All values (CFU/100 mL) |
| BC-CU |
12 |
12 |
47 |
1986 |
2 (17%) |
12, 15, 15, 21, 26, 44, 50, 53, 60, 75, 152, 1986 |
| BC-55 |
13 |
6 |
53 |
866 |
4 (31%) |
6, 20, 23, 24, 26, 28, 53, 75, 131, 145, 166, 378, 866 |
| BC-30 |
3 |
36 |
105 |
517 |
1 (33%) |
36, 105, 517 |
| BC-Can |
8 |
2 |
16 |
30 |
0 (0%) |
2, 3, 5, 7, 16, 17, 27, 30 |
| BC-Eben |
3 |
6 |
28 |
30 |
0 (0%) |
6, 28, 30 |
| All BC |
39 |
2 |
30 |
1986 |
7 (18%) |
median = 30 • mean = 171 • ≥126: 7 of 39 |
Field Colilert Calibration — Boulder Creek
Colilert grab samples matched to Boulder Creek sensor readings (LED 512). E. coli is regressed on the sensor fluorescence, water temperature and turbidity, with a per-sensor fixed effect:
log₁₀(colilert) = b₀ + FE[sensor] + bmon2·mon2 + btemp·temperature + bturb·NTU
NTU = max(0, −145.89 + 2.0488·signal-per-SPAD). The fixed effect (reference 50046) sets each sensor's intercept; the slopes are shared. Source data: ⬇ field_matched_512.csv.
All matched — n=45, R²=0.50, LOO R²=0.21, RMSE=0.49 log₁₀
Post burn-in — n=31, R²=0.49, LOO R²=0.07, RMSE=0.53 log₁₀
Combined Calibration — Lab + Field Colilert + CBT
One pooled model across every reference dataset: lab Colilert (sensors 50030/31/32, controlled dilutions), field Colilert on Boulder Creek, and Aquagenx CBT from the Amazi Meza (Rwanda) and DRIP (Kenya) deployments. E. coli is regressed on the sensor fluorescence and water conditions with a per-sensor fixed effect:
log₁₀(E. coli + 1) = b₀ + FE[sensor] + bmon2·mon2 + btemp·temperature + bturb·NTU + bm×t·mon2·temperature
mon2 is the TLF fluorescence at LED 512; NTU = max(0, −145.89 + 2.0488·signal-per-SPAD); temperature is the in-sensor reading. The mon2×temperature interaction lets the fluorescence slope vary with temperature (it turns positive in warmer water). Each sensor appears under a single reference method, so its fixed effect also carries the offset between methods while the slopes are estimated across all observations. Source data: ⬇ combined_calibration.csv.
n = 446 (295 lab · 45 field · 106 CBT across 50065/50045/50053, 12 sensors) — R² = 0.51, LOO R² = 0.46, RMSE = 0.60 log₁₀. One nearest sensor reading per grab×sensor; temperature is the water temperature every unit reports. Within-source R²: lab 0.74, field 0.35, CBT reads only the censored 0–100 range. 36 CBT samples sit at the Aquagenx 100 CFU detection limit (plotted at the cap).
Binary Detection: ≥126 CFU/100 mL (combined model)
Thresholding the combined model's predicted E. coli at the EPA single-sample recreational limit (126 CFU/100 mL). Left: field Colilert data only. Right: all pooled observations (lab + field + CBT). The ROC sweeps the predicted-concentration threshold; the confusion matrix uses the Youden-optimal cut. (CBT tops out at 100 CFU, so it contributes only negatives.)
Lab Calibration Data
Jan 2–6, 2026 calibration sessions (paper training range). Fluorescence signal (mon2_val) at the paper operating point: led_power = 1024, sipm_bias = 3040. CBT calibration data is in the field-calibration table above.
⬇ Download full dataset (CSV)
Loading lab validation data…
Lab Binary Logistic — ≥126 CFU/100 mL Production binary model
This is the binary alert classifier deployed to all production dashboards (Boulder, Denver, DC). Logistic regression on the full lab dataset (Jan 2–21 2026, n = 300), binary label Colilert ≥ 126 CFU/100 mL. Features: mon2_val_512, floor_temp, tof_mean (z-scored). Performance estimated by leave-one-out cross-validation. Single-event caveat: the 9 positive examples all come from the Jan 8 contamination event (three consecutive 25-min time slots), so when any one positive row is held out the other 8 from the same event remain in training. Positive-class sensitivity is over-optimistic for an unseen new event.
Fitting logistic regression with LOO-CV…
Lab: Predicted vs. Observed — Operating Point Comparison
Pooled OLS (Jan 2–6, n = 125): log₁₀(colilert) ~ barcode + signal × floor_temp × tof_mean. Barcode is a fixed-effect intercept shift (reference: 50030); slopes shared. Fit separately for each LED/bias operating point. In-sample R².
LED = 1024 · bias = 3040 — paper
LED = 512 · bias = 3000 — production
LED = 256 · bias = 3300 — original
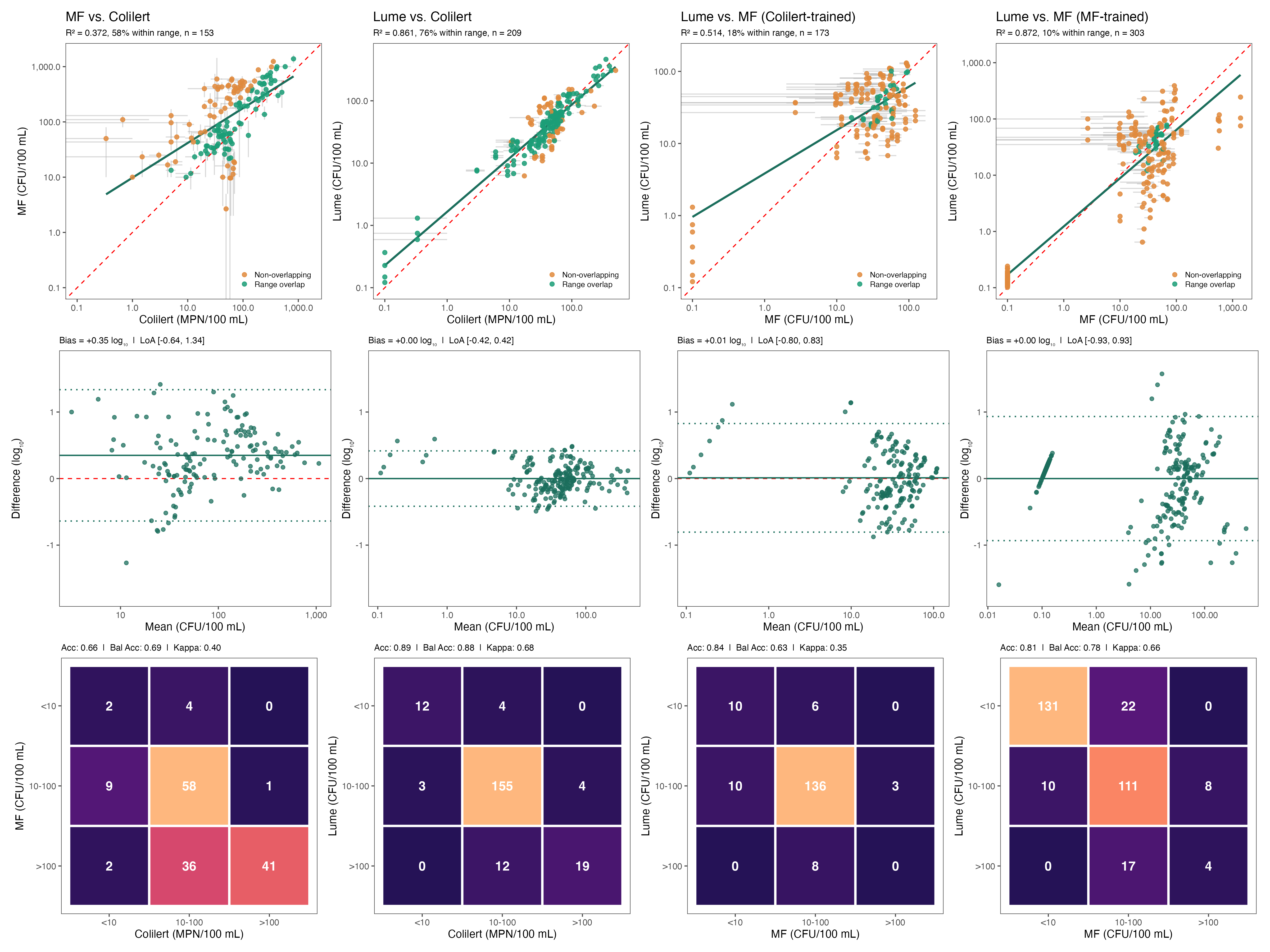
Four-Panel Method Comparison
How does the Lume compare against the two EPA-approved laboratory methods — Colilert (IDEXX) and membrane filtration (MF) — and against itself when retrained on a different reference? Each column analyzes paired samples across three frameworks: log-log regression (top), Bland-Altman agreement (middle), and categorical classification (bottom).
Column 1 · MF vs. Colilert (n = 153)
The dedicated method comparison study pairs Colilert (n = 2 replicates) with membrane filtration (n = 3 replicates) across 161 datetimes; 8 zero-valued pairs are excluded from the log-scale analysis, yielding 153 observations. The two EPA-approved methods show R² = 0.572 with a +0.35 log10 bias — MF systematically reads ~2.2× higher than Colilert. 95% limits of agreement span [−0.64, +1.34], meaning paired lab samples can differ by up to ~22× in either direction. Categorical accuracy is 0.66 (Cohen’s κ = 0.40), i.e. “fair” agreement. This inter-method disagreement sets the ceiling for what any sensor can be expected to achieve against either reference.
Column 2 · Lume vs. Colilert (n = 209, Colilert-trained)
The Colilert-trained Lume regression is evaluated against Colilert across all bench (n = 176) and field (n = 33) observations. The sensor achieves R² = 0.881, a bias of 0.00 log10, and tight limits of agreement [−0.42, +0.42] — Lume predictions stay within ~2.6× of the reference. Categorical accuracy is 0.89 with κ = 0.88, which is “almost perfect” agreement. Against its training reference, the Lume performs as well as or better than the two EPA methods perform against each other.
Column 3 · Lume vs. MF (n = 173, Colilert-trained)
The same Colilert-trained Lume model is now evaluated against membrane filtration — a reference method it was never trained on. Performance drops to R² = 0.514 with LoA [−0.80, +0.83] and categorical accuracy 0.84 (κ = 0.65). Critically, the ~0.37 drop in R² from column 2 to column 3 is of the same order as the inter-method disagreement between Colilert and MF themselves (column 1, R² = 0.572). Most of the apparent loss is attributable to reference-method disagreement, not sensor limitations.
Column 4 · Lume vs. MF (n = 303, MF-trained)
To isolate the effect of reference-method choice, the Lume regression is refit using MF as the training target, over the full bucket dataset. Performance jumps back to R² = 0.872 — essentially matching the Colilert-trained model against Colilert. Bias is 0.00 with LoA [−0.93, +0.93]; the slightly wider LoA reflects the higher within-method variability of MF replicates (57.9% RPD vs. 43.5% for Colilert), not a sensor deficiency. Categorical accuracy is 0.81 (κ = 0.66).
Headline Finding
Sensor-to-reference agreement is bounded by reference-method reproducibility, not by Lume hardware. Whichever culture method is adopted as truth, the Lume fits it at R² ≈ 0.87–0.88. The gap between columns 2 and 3 is almost exactly the disagreement between the two lab methods themselves (column 1). The Lume is method-agnostic; its ceiling is set by the reference it is trained against, and it already achieves quantitative performance at or above the inter-method agreement ceiling between the two accepted laboratory techniques — while providing continuous temporal coverage that grab-sample laboratory methods cannot.